Note
Go to the end to download the full example code.
Mixture Model#
This notebook contains the code for a simple implementation of the Leaspy Mixture model on synthetic data. Before implementing the model take a look at the relevant mathematical framework in the user guide.
The following imports are required libraries for numerical computation, data manipulation, and visualization.
import os
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import torch
import leaspy
from leaspy.io.data import Data, Dataset
This toy example is part of a simulation study, carried out by Sofia Kaisaridi that will be included in an article to be submitted in a biostatistics journal. The dataset contains 1000 individuals each with 6 visits and 6 scores.
leaspy_root = os.path.dirname(leaspy.__file__)
data_path = os.path.join(leaspy_root, "datasets/data/simulated_data_for_mixture.csv")
all_data = pd.read_csv(data_path, sep=";", decimal=",")
all_data["ID"] = all_data["ID"].ffill()
all_data = all_data.set_index(["ID", "TIME"])
all_data.head()
We load the Mixture Model from the leaspy library and transform the dataset in a leaspy-compatible form with the built-in functions.
from leaspy.models import LogisticMultivariateMixtureModel
leaspy_data = Data.from_dataframe(all_data)
leaspy_dataset = Dataset(leaspy_data)
Then we fit a model with 3 clusters and 2 sources. Note that we have an extra argument n_clusters than the standard model that has to be specified in order for the mixture model to run.
model = LogisticMultivariateMixtureModel(
name="multi",
source_dimension=2,
dimension=6,
n_clusters=3,
obs_models="gaussian-diagonal",
)
model.fit(leaspy_dataset, "mcmc_saem", seed=1312, n_iter=100, progress_bar=False)
==> Setting seed to 1312
Fit with `AlgorithmName.FIT_MCMC_SAEM` took: 4s
First we take a look in the population parameters. With the mixture model we obtain separate values for the tau_mean, xi_mean and the sources_mean for each cluster, as well as the cluster probabilities (probs).
print(model.parameters)
{'betas_mean': tensor([[-0.0099, 0.0054],
[-0.0062, 0.0054],
[ 0.0372, -0.0189],
[ 0.0329, -0.0143],
[ 0.0424, -0.0385]]), 'log_g_mean': tensor([ 0.6386, 1.6792, 2.2111, 0.9981, 0.4451, -0.0358]), 'log_v0_mean': tensor([-5.0169, -5.0958, -6.3110, -5.1638, -5.3070, -5.6924]), 'noise_std': tensor([0.0744, 0.0715, 0.0370, 0.0625, 0.0812, 0.0779], dtype=torch.float64), 'probs': tensor([0.3247, 0.4001, 0.2752], dtype=torch.float64), 'sources_mean': tensor([[-1.1791, -0.9338, 2.7506],
[ 2.4279, -0.8996, -1.5589]], dtype=torch.float64), 'tau_mean': tensor([71.0264, 66.0683, 57.4709], dtype=torch.float64), 'tau_std': tensor([10.4716, 8.8100, 11.6153], dtype=torch.float64), 'xi_mean': tensor([-0.1970, -0.2768, 0.6348], dtype=torch.float64), 'xi_std': tensor([0.8801, 0.9058, 1.0671], dtype=torch.float64)}
Then we can also retrieve the individual parameters and the posteriors probabilities of cluster membership. We then consider that the cluster label is the cluster with the biggest probability.
from torch.distributions import Normal
def get_ip(df_leaspy, model):
"""
leaspy_data : the dataframe with the correct indexing
leaspy_mixture : the leaspy object after the fit
"""
ip = pd.DataFrame(df_leaspy.index.get_level_values("ID").unique(), columns=["ID"])
ip[["tau"]] = model.state["tau"]
ip[["xi"]] = model.state["xi"]
ip[["sources_0"]] = model.state["sources"][:, 0].cpu().numpy().reshape(-1, 1)
ip[["sources_1"]] = model.state["sources"][:, 1].cpu().numpy().reshape(-1, 1)
params = model.parameters
probs = params["probs"]
# Number of individuals and clusters
n = len(ip)
k = len(probs)
means = {
"tau": params["tau_mean"], # shape: (2,)
"xi": params["xi_mean"],
"sources_0": params["sources_mean"][0, :],
"sources_1": params["sources_mean"][1, :],
}
stds = {
"tau": params["tau_std"],
"xi": params["xi_std"],
"sources_0": torch.tensor(1),
"sources_1": torch.tensor(1),
}
stds["sources_0"] = stds["sources_0"].repeat(k)
stds["sources_1"] = stds["sources_1"].repeat(k)
values = {
"tau": torch.tensor(ip["tau"].values),
"xi": torch.tensor(ip["xi"].values),
"sources_0": torch.tensor(ip["sources_0"].values),
"sources_1": torch.tensor(ip["sources_1"].values),
}
# Compute log-likelihoods for each variable
log_likelihoods = torch.zeros((n, k))
for var in ["tau", "xi", "sources_0", "sources_1"]:
x = torch.tensor(ip[var].values)
for cluster in range(k):
dist = Normal(means[var][cluster], stds[var][cluster])
log_likelihoods[:, cluster] += dist.log_prob(x)
# Add log-priors
log_priors = torch.log(probs)
log_posteriors = log_likelihoods + log_priors
# Normalize using logsumexp
log_sum = torch.logsumexp(log_posteriors, dim=1, keepdim=True)
responsibilities = torch.exp(log_posteriors - log_sum)
for i in range(responsibilities.shape[1]):
ip[f"prob_cluster_{i}"] = responsibilities[:, i].numpy()
# Automatically find all probability columns
prob_cols = [col for col in ip.columns if col.startswith("prob_cluster_")]
# Assign the most likely cluster
ip["cluster_label"] = ip[prob_cols].values.argmax(axis=1)
return ip
ip = get_ip(all_data, model)
ip.head()
We produce the population progression plots. We separate the model parameters for each cluster and we store them to a dedicated item.
from leaspy.io.outputs import IndividualParameters
parameters = model.parameters
number_of_sources = 2
# cluster 0
mean_xi = parameters["xi_mean"].numpy()[0]
mean_tau = parameters["tau_mean"].numpy()[0]
mean_source = parameters["sources_mean"].numpy()[0, 0].tolist()
mean_sources = [mean_source] * number_of_sources
parameters_0 = {"xi": mean_xi, "tau": mean_tau, "sources": mean_sources}
ip_0 = IndividualParameters()
ip_0.add_individual_parameters("average", parameters_0)
# cluster 1
mean_xi = parameters["xi_mean"].numpy()[1]
mean_tau = parameters["tau_mean"].numpy()[1]
mean_source = parameters["sources_mean"].numpy()[0, 1].tolist()
mean_sources = [mean_source] * number_of_sources
parameters_1 = {"xi": mean_xi, "tau": mean_tau, "sources": mean_sources}
ip_1 = IndividualParameters()
ip_1.add_individual_parameters("average", parameters_1)
# cluster 2
mean_xi = parameters["xi_mean"].numpy()[2]
mean_tau = parameters["tau_mean"].numpy()[2]
mean_source = parameters["sources_mean"].numpy()[0, 2].tolist()
number_of_sources = 2
mean_sources = [mean_source] * number_of_sources
parameters_2 = {"xi": mean_xi, "tau": mean_tau, "sources": mean_sources}
ip_2 = IndividualParameters()
ip_2.add_individual_parameters("average", parameters_2)
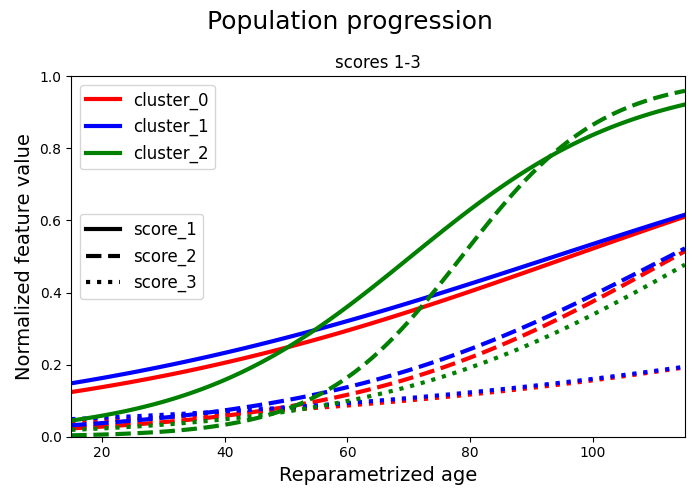
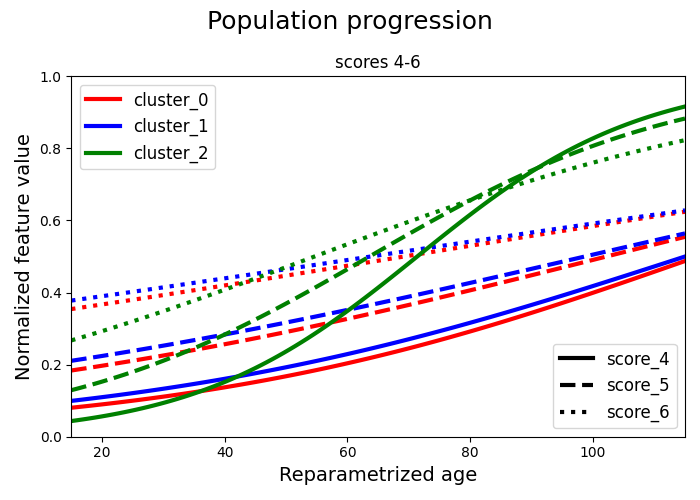
We separate the scores and we plot two graphs for clarity. Each cluster is represented with a different colour. Each score is represented with a different linestyle. In the first graph we have the first 3 scores and in the second graph we have the remaining 3 scores. In each graph we plot all the clusters.
from matplotlib.lines import Line2D
timepoints = np.linspace(15, 115, 105)
values_0 = model.estimate({"average": timepoints}, ip_0)
values_1 = model.estimate({"average": timepoints}, ip_1)
values_2 = model.estimate({"average": timepoints}, ip_2)
# A. Scores 1-3
plt.figure(figsize=(7, 5))
plt.ylim(0, 1)
lines = [
"-",
"--",
":",
]
## cluster 0
for ls, name, val in zip(lines, model.features, values_0["average"][:, 0:3].T):
plt.plot(timepoints, val, label=f"cluster_0_{name}", c="red", linewidth=3, ls=ls)
## cluster 1
for ls, name, val in zip(lines, model.features, values_1["average"][:, 0:3].T):
plt.plot(timepoints, val, label=f"cluster_1_{name}", c="blue", linewidth=3, ls=ls)
## cluster 2
for ls, name, val in zip(lines, model.features, values_2["average"][:, 0:3].T):
plt.plot(timepoints, val, label=f"cluster_2_{name}", c="green", linewidth=3, ls=ls)
## legends
cluster_legend = [
Line2D([0], [0], color="red", linewidth=3, label="cluster_0"),
Line2D([0], [0], color="blue", linewidth=3, label="cluster_1"),
Line2D([0], [0], color="green", linewidth=3, label="cluster_2"),
]
legend1 = plt.legend(handles=cluster_legend, loc="upper left", prop={"size": 12})
color_legend = [
Line2D([0], [0], linestyle=lines[0], color="black", linewidth=3, label="score_1"),
Line2D([0], [0], linestyle=lines[1], color="black", linewidth=3, label="score_2"),
Line2D([0], [0], linestyle=lines[2], color="black", linewidth=3, label="score_3"),
]
legend2 = plt.legend(
handles=color_legend, loc="center left", title_fontsize=14, prop={"size": 12}
)
plt.gca().add_artist(legend1)
plt.xlim(min(timepoints), max(timepoints))
plt.xlabel("Reparametrized age", fontsize=14)
plt.ylabel("Normalized feature value", fontsize=14)
plt.title("scores 1-3")
plt.suptitle("Population progression", fontsize=18)
plt.tight_layout()
plt.show()
# B. Scores 4-6
plt.figure(figsize=(7, 5))
plt.ylim(0, 1)
lines = [
"-",
"--",
":",
]
## cluster 0
for ls, name, val in zip(lines, model.features, values_0["average"][:, 3:6].T):
plt.plot(timepoints, val, label=f"cluster_0_{name}", c="red", linewidth=3, ls=ls)
## cluster 1
for ls, name, val in zip(lines, model.features, values_1["average"][:, 3:6].T):
plt.plot(timepoints, val, label=f"cluster_1_{name}", c="blue", linewidth=3, ls=ls)
## cluster 2
for ls, name, val in zip(lines, model.features, values_2["average"][:, 3:6].T):
plt.plot(timepoints, val, label=f"cluster_2_{name}", c="green", linewidth=3, ls=ls)
## legends
cluster_legend = [
Line2D([0], [0], color="red", linewidth=3, label="cluster_0"),
Line2D([0], [0], color="blue", linewidth=3, label="cluster_1"),
Line2D([0], [0], color="green", linewidth=3, label="cluster_2"),
]
legend1 = plt.legend(handles=cluster_legend, loc="upper left", prop={"size": 12})
color_legend = [
Line2D([0], [0], linestyle=lines[0], color="black", linewidth=3, label="score_4"),
Line2D([0], [0], linestyle=lines[1], color="black", linewidth=3, label="score_5"),
Line2D([0], [0], linestyle=lines[2], color="black", linewidth=3, label="score_6"),
]
legend2 = plt.legend(
handles=color_legend, loc="lower right", title_fontsize=14, prop={"size": 12}
)
plt.gca().add_artist(legend1)
plt.xlim(min(timepoints), max(timepoints))
plt.xlabel("Reparametrized age", fontsize=14)
plt.ylabel("Normalized feature value", fontsize=14)
plt.title("scores 4-6")
plt.suptitle("Population progression", fontsize=18)
plt.tight_layout()
plt.show()
Total running time of the script: (0 minutes 5.356 seconds)